What are Phages?
A phage (an abbreviated term for a bacteriophage) is a virus that infects bacteria. They are made up of of a nucleic acid molecule that is surrounded by a protein structure. Phages produce viral components by first hijacking the cellular machinery of a bacterium. These viral components are then assembled to form new phages. When the phage particles are released through a process called lysis, the bacterium cell is destroyed. Since phages typically have a relatively simple genome and multiply rapidly, they have been exploited as a useful technique known as phage display.
Uses of Phage Display
Phage display was first described in 1985 when foreign DNA fragments were inserted into a gene encoding a coat protein of filamentous phage, a type of bacteriophage which infects E. coli. Without impacting on the phage’s ability to infect the bacteria, the resulting fusion protein was displayed at the phage surface. It is possible to enrich protein-expressing phages from the supernatant of bacterial cultures by using antibodies directed against the foreign component of the protein. This approach directly linked the genotype of the phage to the phenotype. This resulted in phage display becoming a preferred method of cloning a gene when an antibody against the gene product is available.
Since its discovery, phage display has become a widely used method to create libraries containing numbers of different protein variants. These can be screened within a very short timescale to provide detailed information regarding the interaction of phage-expressed proteins with specific binding partners. To achieve this, the phage display library is exposed to a target molecule which has been immobilized on a solid support. Following incubation, unbound phages are washed away, and bound phages eluted. Phages with specificity for the target can be used to infect new host cells for amplification. Alternatively, multiple rounds of incubation, washing and elution can be performed to select phages which bind with the greatest efficiency; this is a process known as panning.
The phage display cycle.
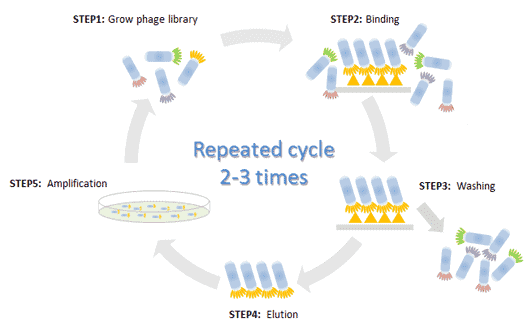
The Phage Display Cycle (By Cusabio).
Although phage display sounds simple, in practice it is highly complex. Consideration must be given to the selection of an appropriate phage display system. This includes the choice of strain, the phage coat protein within which the foreign DNA will be inserted, and whether a helper phage is required to supply essential structural proteins. It is also necessary to consider the type of support on which the target will be immobilised (e.g. microtiter plate wells, PVDF membrane, column matrix, magnetic beads) and to carefully optimise each phase of the selection cycle. In addition, a well-designed panning procedure is essential to identify phages most suitable to the ensuing downstream application.
By partnering with an experienced provider of phage display services, researchers can receive different benefit. Wether it is from high-quality phage display library construction or custom phage display library screening services. The technology can save significant amounts of time and resource, while providing unique insight to accelerate research. At Pivotal Scientific, we’re well placed to introduce you to experts in phage display, whatever your research requirements are.